1
2
3
4
5
6
7
8
9
10
11
12
13
14
15
16
17
18
19
20
21
22
23
24
25
26
27
28
29
30
31
32
|
readr::read_delim("./pi-per-site-acuce_windows-50000.csv", col_names = FALSE) %>%
magrittr::set_names(c("chr", "pos", "pi")) %>%
dplyr::mutate(chr = stringr::str_replace(chr, "chr", "") %>% as.numeric(),
snp = paste0(chr, pos))-> df.pi
df.pi %>%
dplyr::mutate(temp = paste0(chr, pi)) %>%
dplyr::group_by(chr) %>%
dplyr::mutate(chr.len = max(pos)) %>%
dplyr::ungroup() %>%
dplyr::mutate(total = cumsum(chr.len) - chr.len) %>%
dplyr::arrange(chr, pos) %>%
dplyr::mutate(cum = pos + total) %>%
dplyr::mutate(new.pi = -log2(pi)) %>%
dplyr::mutate(rs = paste0(chr, pos)) %>%
dplyr::mutate(chr = stringr::str_replace(chr, "chr", "") %>% as.numeric())-> df
df %>%
group_by(chr) %>%
summarize(center=(max(cum) + min(cum) ) / 2) -> X_axis
df %>%
ggplot(aes(cum, pi)) +
geom_jitter(aes(color = as.factor(chr))) +
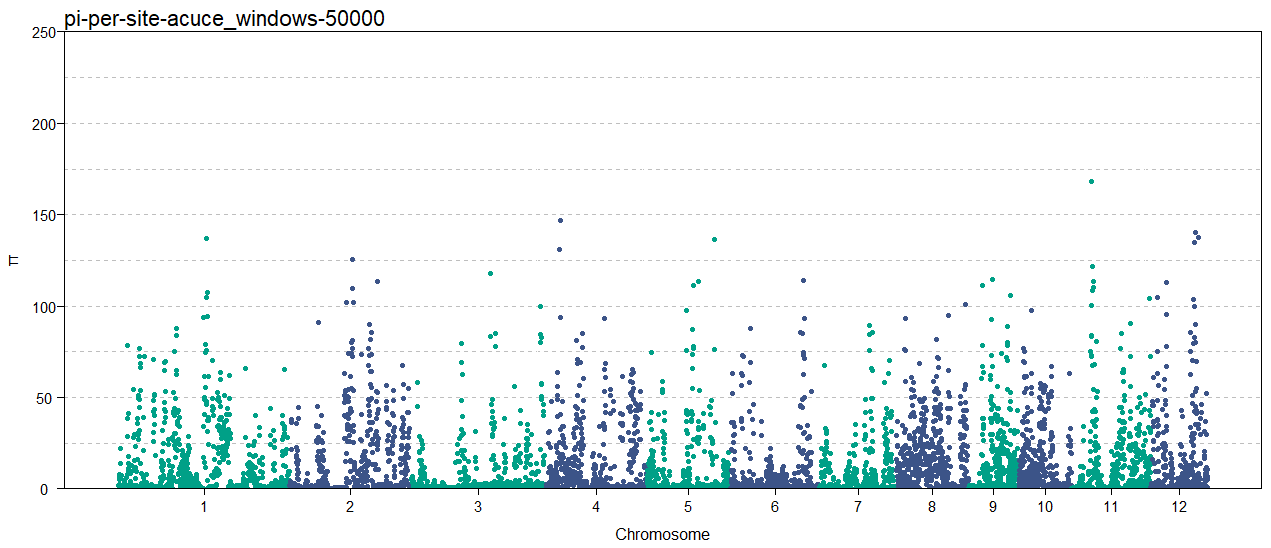
labs(x = "Chromosome", y = "π", title = "pi-per-site-acuce_windows-50000") +
scale_color_manual(values = rep(c("#00A087FF", "#3C5488FF"), 6)) +
scale_x_continuous( label = X_axis$chr, breaks= X_axis$center) +
scale_y_continuous(expand = c(0, 0), limits = c(0,250), breaks = seq(0, 250,50)) +
pac4xiang::mytheme_cn() +
theme(legend.position = "none") -> p.3a.1
|